Introduction
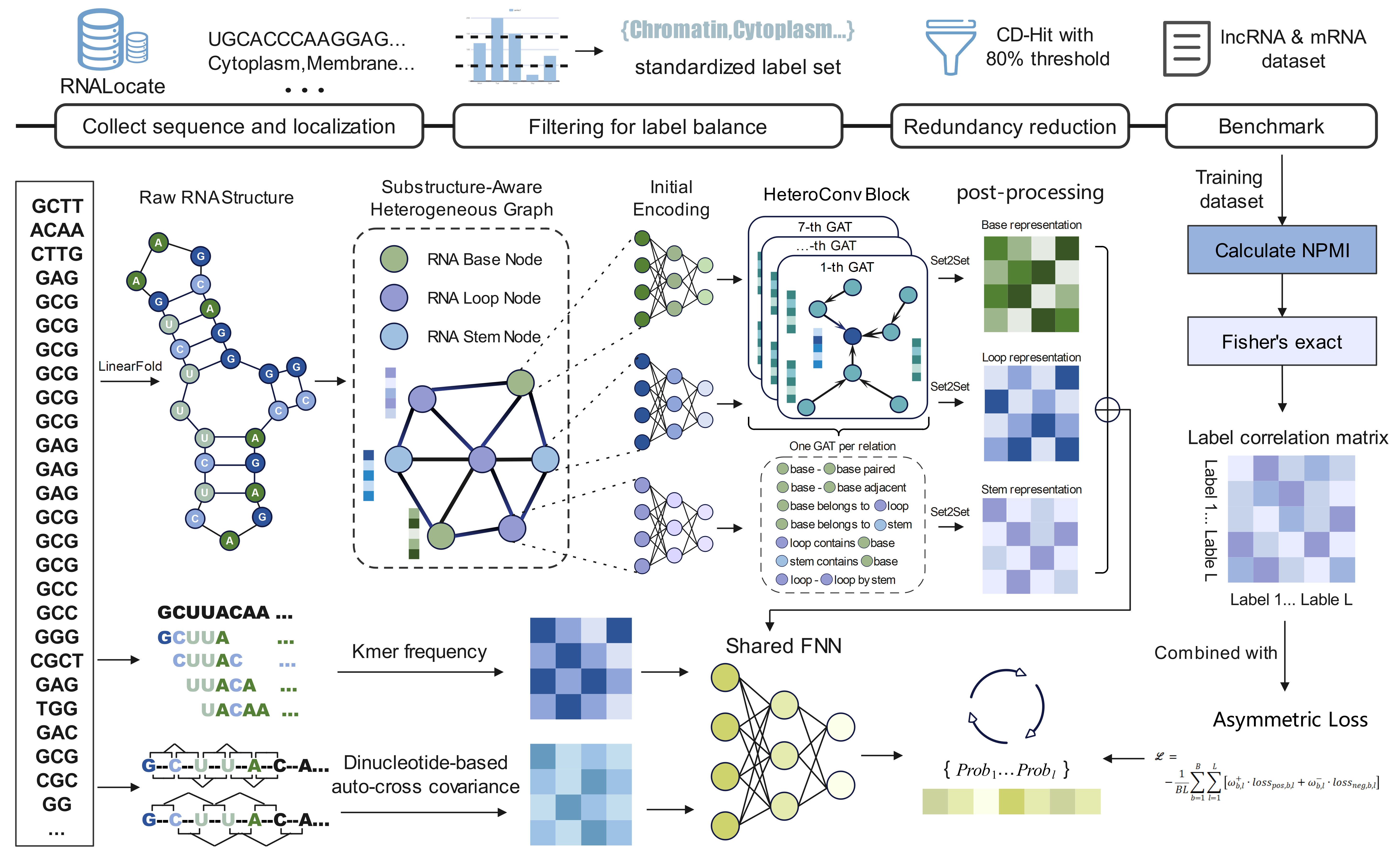
We propose a unified graph neural network-based framework for RNA subcellular localization prediction, named GRASP, which is applicable to both lncRNAs and mRNAs. The framework adopts an RNA substructure-aware heterogeneous graph modeling strategy: RNA molecules are represented using nucleotide nodes together with secondary-structure-derived substructure nodes (e.g. loops and stems) and their associated relational edges, enabling joint modeling of base-level interactions and regional structural context for multi-label localization prediction.